Portfolio
Simulation tools, numerical methods, reproducible workflows.

|
Chaste: Cancer, Heart and Soft Tissue Environment Collaborators: Universities of Sheffield and Nottingham Funder: BBSRC |
Chaste is a C++ simulation package with Python bindings, designed for computationally intensive problems in biology and physiology. It supports tissue- and cell-level electrophysiology, discrete and agent-based tissue modelling, and soft tissue modelling. Full project page here. |
|
POSEIDON: A model of fish, fishers, and fisheries. Collaborators: Prof. Richard Bailey, School of Geography and the Environment Funder: UK Research and Innovation (UKRI) |
POSEIDON is an advanced simulation tool designed to help manage fisheries by modelling the behaviour of fishing fleets and their interactions with the environment. Full project page here. |
|
|
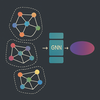
Local2Global: Infer global embeddings from local graph embeddings Collaborators: Mathematics Institute, University of Warwick Funder: Turing Institute |
A Python library for building patch-based graph structures, including temporal graphs, using Graph Neural Networks. It infers global embeddings from local patches for tasks like anomaly detection, event prediction, and graph-level classification. |
|
PyBaMM: An open-source battery simulation package written in Python. Collaborators: Mathematical Institute - Prof Jon Chapman and Prof Colin Please Funder: The Faraday Institution |
PyBaMM consists of (i) a framework for writing and solving systems of differential equations, (ii) a library of battery models and parameters, and (iii) specialized tools for simulating battery-specific experiments and visualizing the results. Full project page here. |
|

|
RoughPy: Streaming data as rough paths Python toolbox Collaborators: DataSig, Mathematical Institute - Prof. Terry Lyons and Dr. Sam Morley Funder: EPSRC, Alan Turing Institute, The Data Centric Engineering Programme |
RoughPy is a package for working with streaming data as rough paths, and working with algebraic objects such as free tensors, shuffle tensors, and elements of the free Lie algebra. |

|
diffsol: High-performance ODE & DAE solvers in Rust & Python |
diffsol is a Rust crate for solving ODEs and semi-explicit DAEs with adaptive, stiff and non-stiff solvers, sensitivity analysis, and event handling. With the DiffSL DSL and pydiffsol bindings, it delivers Rust-level performance to Python users |

|
RELION: REgularised LIkelihood OptimisatioN Collaborators: Scientific Computing, Science and Technology Facilities, Harwell Science & Innovation Campus Funder: Scientific Computing, Science and Technology Facilities |
RELION employs empirical Bayes for electron cryo-microscopy structure determination. OxRSE have assisted with code optimisation and GPU use, leading to improved computational efficiency. |
Infrastructure for data collection, processing & visualisation.
|
Global.health: Epidemiological data platform for tracking infectious diseases Collaborators: Prof. Moritz Kraemer, Department of Biology Funder: Google.org, Oxford Martin School, Rockefeller, Wellcome Trust |
Global.health is an epidemiological platform established during the COVID-19 pandemic to provide open access to de-identified line list case data, facilitating research into infectious diseases and emerging outbreaks. The platform’s primary mission is to deliver timely and accurate data to decision-makers, researchers, and the public during the critical first 100 days of an outbreak. Full project page here. |
|

|
OxWell: Interactive tool for exploring youth mental health survey data Collaborators: Department of Psychiatry |
The OxWell data explorer is an interactive platform to support exploration of mental health survey data from children and young people, enabling researchers and policymakers to visualise trends and inform evidence-based decisions. Full project page here. |

|
GRAPEVNE: Graphical Analytical Pipeline Development Environment Collaborators: Pandemic Sciences Institute, Department of Biology |
GRAPEVNE is an interactive environment for building and validating data processing workflows built around the Snakemake workflow manager. Originally focussed on solving phylogenetic problems, GRAPEVNE has developed to support a broad range of use-cases. |

|
InsightBoard: An AI-assisted dashboard for managing line-list data. Collaborators: Pandemic Sciences Institute, Department of Biology |
InsightBoard is a powerful tool for ingesting and reporting on data from multiple sources. Seamlessly transform various data formats into a unified schema, store them in a central database, and generate custom reports, including interactive visualizations. |

|
Galv: Data and metadata management platform for battery science Collaborators: Battery Intelligence Lab |
The Galv project is a data and metadata management platform for battery science. Labs conducting battery research can use the platform to store and share (meta)data created from experimental electrochemical experiments. |
|
REF 2021 Data Explorer: Explore, analyse, and visualise REF 2021 research data |
The REF 2021 Data Explorer enables users to search, filter, visualise, and download processed data from the UK’s 2021 Research Excellence Framework. It supports in-depth analysis of outputs, impact, research income, and environment across institutions. Full project page here. |
|

|
DART-Pipeline: Dengue Advanced Readiness Tools Collaborators: OeRC, Department of Biology, OUCRU Funder: Wellcome Trust |
DART (Dengue Advanced Readiness Tools) is a dengue outbreak forecasting system by OUCRU, funded by Wellcome Trust. Its goal is to offer high-resolution, real-time to 4-week forecasts at district level, starting in Ho Chi Minh City and Hanoi, to aid targeted responses. |
| ISARIC: International Severe Acute Respiratory and emerging Infection Consortium
Collaborators: Nuffield Department of Medicine, Pandemic Sciences Institute Funder: Wellcome Trust |
ISARIC is a global federation of 60+ clinical research networks conducting agile, coordinated research on infectious diseases like COVID-19, Ebola, dengue, and more to improve patient care and global outbreak response. Full project page here. |
|

|
GLACIER: Graphical Launchpad for Analysis, Computation, Inference and Explication of Results Collaborators: ARTIC network, University of Edinburgh Funder: global.Health |
GLACIER is a genreal purpose platform for provisioning, launching, monitoring and reporting on analytic workflows. Our current use-case focusses on bioinformatic and genomic applications. |
Platforms for research engagement, collaboration, and delivery.
|
LanguageScreen: App-based screening of children’s early language skills Collaborators: Department of Education & Department of Experimental Psychology |
LanguageScreen is a mobile app developed with education researchers to assess young children’s language skills. Used in UK and worldwide, it supports early language interventions in schools and led to the OxEd & Assessment spin-out. Full project page here. |
|
|
MAMA: A mobile app for MAMA Study participants Collaborators: NPEU Clinical Trials Unit Funder: NIHR Health Technology Assessment (HTA) Programme |
The MAMA Trial app supports participants in the NIHR-funded MAMA clinical study by enabling easy access to trial information and questionnaires. Developed with NPEU CTU, it streamlines participation and enhances user engagement. |
|
PETRUSHKA: Personalising Antidepressant Treatment for Unipolar Depression Collaborators: Department of Psychiatry |
Personalising Antidepressant Treatment for Unipolar Depression Combining Individual Choices, Risks and big Data Full project page here. |
|
|
OCSPlus: App-based cognitive screening for mild cognitive impairment Collaborators: Department of Experimental Psychology, Nuffield Department of Clinical Neuroscience |
The Oxford Cognitive Screen-Plus (OCS-Plus) is a mobile app-based tool developed to sensitively detect domain-general cognitive impairments, including memory and executive function, in conditions like stroke, MCI, and ageing, without overloading language demands. Full project page here. |
|
| Oxford Multiple Errands Task:
App-based tool for cognitive screening for executive dysfunction Collaborators: Department of Experimental Psychology, Nuffield Department of Clinical Neuroscience |
The Oxford Multiple Errands Test (OxMET) is a brief, tablet-based screening tool that simulates everyday shopping to assess executive functions like planning and goal management, with high accessibility for stroke survivors and strong real-life relevance. Full project page here. |
|

|
PKPD Explorer: User-friendly web app for exploration of PK/PD models Funder: Roche |
PKPD Explorer is an open source web-based application, enabling both modellers and non-modellers to explore, share, analyse and model the pharmacokinetics and pharmacodynamics of chemical compounds. |

|
Schoolpol: The Transformation of Post War Education: Causes & Effects Collaborators: Department of Politics and International Relations, Department of Social Policy and Intervention Funder: European Research Council |
Schoolpol examines the long-term impact of education on social and political outcomes and provides crucial resources for policymakers and help support educational reform. |

|
EDC: Epilepsy Diagnostic Companion Collaborators: Nuffield Department of Cognitive Neuroscience, Center for Global Epilepsy |
A mobile app designed to assist healthcare workers in diagnosing convulsive epileptic seizures, especially in resource-limited settings. |

|
Gutenberg: An open source research and teaching platform Collaborators: University of Edinburgh, Imperial University, University of Southampton Funder: SPF ExCALIBUR programme |
Provides an environment for hosting training materials in a structured, pathway-based style, offering a visual interface for browsing the various sections of training content and viewing them. Full project page here. |
|
SPIRES: Real-time MAMS-capable pathway assignment Collaborators: Precision Psychiatry Lab |
SPIRES allows researchers to easily build powerful APIs that assign participants to treatment paths responsively and in real-time. Designed with MAMS trials in mind. |

|
Bushel: A bulk uploader for FigShare resources Collaborators: Sustainable Digital Scholarship (SDS) Service |
The Bushel tool enables the Sustainable Digital Scholarship team to batch-upload resources to the FigShare data archive. |

|
Jamaica Systemic Resilience Assessment Tool: Climate-related risk analytics for infrastructure in Jamaica Collaborators: Oxford Programme for Sustainable Infrastructure Systems |
The Jamaica Systemic Risk Assessment Tool (J‑SRAT) supports climate adaptation decision-making by identifying spatial criticalities and risks under current and future climate scenarios. |

|
OpenBES: Simulate and monitor building energy requirements and usage. Collaborators: Department of Engineering Science, ZERO Institute, University of Edinburgh Funder: EPSRC (IAA 2025-6) |
OpenBES (Open Building Energy Simulation) is a engineering survey tool that produces detailed energy requirement and usage data from building and climate information. Full project page here. |
Research software engineering support for the Schmidt AI in Science programme's fellows.
|
Schmidt AI Fellowship Support: Dedicated RSE support for Schmidt AI in Science Fellowships Funder: Schmidt Sciences |
Three full-time RSEs provide support for postdoctoral fellows with help on individual projects as well as training in software best practices, helping fellows build robust, reusable tools and gain valuable, transferable skills for future careers. Full project page here. |
|

|
Bee Behaviour Tracking: Analysing behaviour changes in bees using movement tracking Collaborators: Dr. Rachel Parkinson - Department of Biology Funder: Schmidt Sciences |
A deep learning system to track bees in experimental chambers in order to analyse how they move and interact with one another. The research aim is to analyse the effects of pesticides on their behaviour. |
|
SpeedyWeather: Play atmospheric modelling like it's LEGO Collaborators: Milan Klöwer, Atmospheric Physics Funder: Schmidt Sciences |
SpeedyWeather.jl is a global atmospheric model with simple physics developed as a research playground with an everything-flexible attitude as long as it is speedy. It is easy to use and easy to extend, making atmospheric modelling an interactive experience. Full project page here. |













